In a previous discussion [What’s wrong with this figure?], I noted that certain things seem to happen with disturbing frequency in discussions of genome size. The first is the invocation of pre-Darwinian “Great Chain of Being” thinking, in which humans are considered the most complex organisms, with all others ranked at lower positions on the scala naturae. Of course, this is not restricted to genomics — one can find references to “lower vertebrates”, “subhuman primates”, or “higher plants” peppered throughout the scientific literature. The second issue is the exclusive use of genome sequence data in discussions of genome size diversity. This is problematic because, with few exceptions, sequencing targets are selected in large part on the basis of having small and manageable genomes. I receive many requests from colleagues to provide genome size estimates, and the hope is always that they will turn out to be small such that they will have a chance of being adopted as a sequencing model. There are obvious pragmatic reasons for this, but it means that one must be careful about interpreting data from an inherently biased set of data.
The previous discussion focused on examples in which authors have tried to demonstrate a link between the amount of non-coding DNA and organismal complexity, by making both of the mistakes outlined above. In this post, I want to discuss the opposite but equally aggravating problem, which is using these same limited data to demonstrate an association between genome size and gene number.
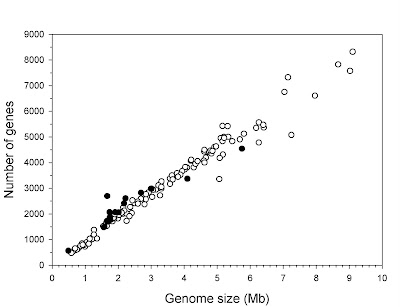
Every now and then, an author makes the claim that gene number and genome size actually are correlated, despite this having been rejected decades ago when the first broad comparisons of genome size were made and the various sorts of non-coding DNA were discovered. The most recent example comes from Lynch (2006):
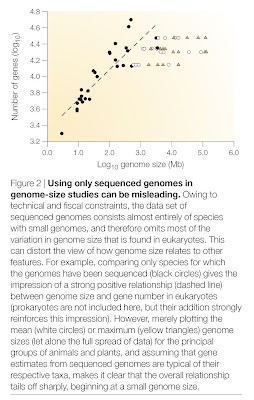
The second problem is, obviously, that this is based on a selective set of species. An estimate of gene number is best achieved with a genome sequence, but genome sequences typically are available only for small genomes. If one assumes that most species in a given group (say, a phylum) have roughly similar gene numbers and plots the actual diversity of genome size (e.g., mean for that phylum), the relationship is nowhere near as clear. Indeed, it drops off completely.
In fact, you can see this happening already in Lynch’s (2006, 2007) figure. Note that there is a totally flat line for the animal data, even though these come from species with comparatively modest genome sizes. Since I work on animals (whose genome sizes range 3,300-fold), I would say that there is no relationship between genome size and gene number in my group. If you compare animals to bacteria, then there is such a relationship, of course, but that almost goes without saying, and could relate to differences in chromosome structure as much as anything else.
The point is that genome sequencing data are extremely useful, including in discussions of genome size, but that they, like all data, must be interpreted within their proper context. Genome sequencing models, at least at the moment, do not encompass the diversity that exists among eukaryotes. In fact, even with 10,000 species in the various databases [animals, plants, fungi], the current dataset of eukaryotic genome size diversity itself is far from comprehensive.
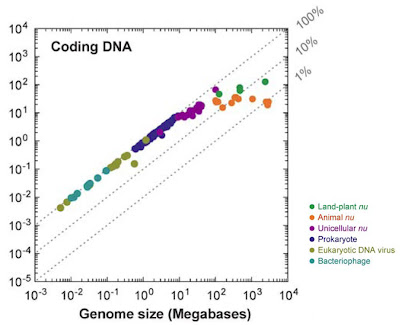
 The diversity of archaeal, bacterial, and eukaryotic genome sizes as currently known from more than 10,000 estimates. From Gregory (2005). Click for larger view.
The diversity of archaeal, bacterial, and eukaryotic genome sizes as currently known from more than 10,000 estimates. From Gregory (2005). Click for larger view.What is clear, and has been for decades, is that genome size evolves independently of organismal complexity and gene number (which themselves may evolve more or less independently of one another). This makes it a very intriguing puzzle to study, one that has resisted all attempts at one-dimensional explanation for over half a century.
___________
References
Gregory, T.R. 2005. Synergy between sequence and size in large-scale genomics. Nature Reviews Genetics 6: 699-708.
Gregory, T.R. and DeSalle, R. 2005. Comparative genomics in prokaryotes. In: The Evolution of the Genome, edited by T.R. Gregory, pp. 585-675. Elsevier, San Diego, CA.
Lynch, M. 2006. Streamlining and simplification of microbial genome architecture. Annual Review of Microbiology 60: 327-349.
Lynch, M. 2007. The Origins of Genome Architecture. Sinauer Associates, Sunderland, MA.
See also
- Genome Size, Complexity, and the C-Value Paradox (Sandwalk)
- Ladders and cranes everywhere! (Pharyngula)

Hello. I want to know, what is the expect in gen size, with the advance of science and sequencing we need best program of the computation to compare genes more big in size?
How much in bp is the gen biggest?
Thanks
Claudia
My mail is oropendola_iris@yahoo.com