Whereas the possibility that non-coding DNA is functional has been a topic of discussion for decades, it recently has come to the fore with the availability of several sequenced genomes which allow signs of function to be detected at the DNA level. The multi-million-dollar ENCODE project is the largest initiative focused on identifying functional elements in the human genome, but many smaller projects are also ongoing in other species such as mice, Drosophila, and other eukaryotes (e.g., Siepel et al. 2005).
For the most part, the way that potentially functional elements are highlighted is by finding regions of the genome that are essentially unchanged among species whose lineages have been separated for very long periods of time. No change in the sequences suggests that they have been preserved in their present state by natural selection — that is, individuals with mutations in these regions were less fit, and only those with no such changes have left an unbroken line of descendants to the present day. A recent analysis by Katzman et al. (2007) in Science indicated that indeed these “ultraconserved” regions are “ultraselected” in the human genome. Because natural selection is the result of differential survival and reproduction due to heritable phenotypic differences, this provides strong evidence that these regions have some important effect — in fact, probably a function — on the organisms carrying them.
It is important to note that elements exhibiting signs of selective constraint make up a small fraction of the total genome of organisms like mammals, on the order of 5%. Ultraconserved elements in particular represent a very tiny portion of the total DNA. It would therefore be a major exaggeration to assume that the demonstration of such sequences implies that all non-coding DNA is functional. Most or all of it might serve a function, but there is no evidence to support this notion at present. It is also inaccurate to suggest that the discovery of some function in non-coding DNA is a total surprise. Even the early proponents of the “selfish DNA” view of non-coding DNA evolution proposed that some elements would end up being functional, most notably in gene regulation. This certainly appears to have been borne out, and it is quite plausible that more than just the ultraconserved elements are involved in the regulation of coding genes.
However, amidst this backdrop of increasingly refined tabulations of conserved elements in animal genomes there are some observations that raise doubts about just how important they are for organismal fitness. In 2004, for example, Marcelo Nóbrega and colleagues put the importance of conserved non-coding DNA to the test — by deleting some of it. Specifically, they removed two fragments of conserved DNA totaling 1,511 kilobases and 845kb in mice and observed the consequences. Or, more accurately, the lack of consequences. In their experiment, the deletion of more than 2 million base pairs of conserved DNA from the mouse genome had no identifiable effects on the development, physiology, or reproduction of the subjects.
Of course, the mice were kept in lab conditions, and it was argued by some that this may be an unrealistic test given that conditions in the wild are much harsher and any detriment to growth or survival may be hidden in the lab.
In the September 2007 issue of the open access journal PLoS Biology, Nadav Ahituv and coworkers report on a similar but more telling experiment, again using deletions of ultraconserved DNA elements in mice. In this case, the authors deleted four elements ranging in length from 222 to 731bp in ultraconserved regions that are invariant among humans, mice, and rats. More importantly, these regions are known to be located in close proximity to genes for which loss of function mutations result in severe abnormalities.
The assumption, therefore, was that if these regions are conserved because they regulate nearby genes, then their removal should disrupt gene function and result in inviable mice. What did they find?
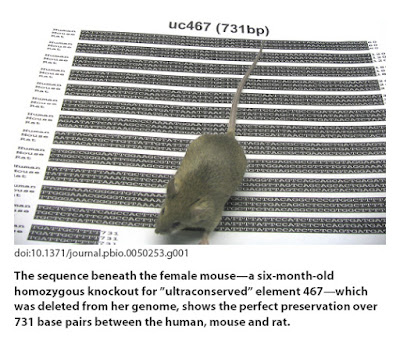 Nothing. No effect whatsoever was detectable in terms of growth, morphology, reproduction, metabolism, or longevity when any of the four elements was deleted. Again, it is possible that some deleterious effect would show up in the wild, or that there is redundancy that allows other elements to regulate these genes if need be, but as far as the expected phenotypic consequences of disrupting the nearby genes goes, it makes no difference whether these specific conserved sequences are present or not.
Nothing. No effect whatsoever was detectable in terms of growth, morphology, reproduction, metabolism, or longevity when any of the four elements was deleted. Again, it is possible that some deleterious effect would show up in the wild, or that there is redundancy that allows other elements to regulate these genes if need be, but as far as the expected phenotypic consequences of disrupting the nearby genes goes, it makes no difference whether these specific conserved sequences are present or not.
At the moment, there is no conclusive evidence one way or another as to the function of most non-coding DNA. It bears noting, however, that although it is very difficult to demonstrate that so much non-coding DNA is non-functional (as this is roughly akin to proving a universal negative), there are reasons to adopt this as the default hypothesis. For example, several mechanisms are known that can generate large amounts of non-coding DNA independent of organismal functions. On the other hand, the evidence for function is thus far restricted to a few percent of the genome, and even here it appears that some of these elements can be eliminated without obvious consequences.
This is not to say that non-coding DNA has no effect; it clearly influences cell size and cell division rate, for example. It is, however, far outstripping the available evidence, and contradicting much of what is already known about genome evolution, to argue that comparative genomics is revealing functions for non-coding DNA at large. At most, genomic analysis is showing genome form, function, and evolution to be much too complex to support any inflexible assumptions on either side.
___________
References
Ahituv, N. et al. (2007). Deletion of ultraconserved elements yields viable mice. PLoS Biology 5(9): e234.
ENCODE Project Consortium (2007). Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447: 799-816.
Gross, L. (2007). Are “ultraconserved” genetic elements really indispensable?. PLoS Biology 5(9): e23.
Katzman, S. et al. (2007). Human genome ultraconserved elements are ultraselected. Science 317: 915.
Nobrega, M.A. et al. (2004). Megabase deletions of gene deserts result in viable mice. Nature 431: 988-993.
Siepal, A. et al. (2005). Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Research 15: 1034-1050.
___________
Update:
Larry Moran has a nice piece on this at Sandwalk. There is also a post about it at This Week in Evolution. Kay of Suicyte (great title — he works on apoptosis) has an interesting post as well. And for goodness sake, could someone please go read Chris Harrison’s earlier post on Interrogating Nature so he can’t stop the crank-like self promotion in all the discussion threads? (Just kidding, Chris –it’s a nice post).

I’m a little confused as to what “ultraconserved” means. Is there evidence that mutations occur in these regions at the same rate as they do in other stretches of the genome?
Anonymous,
Mutations occur at roughly the same rate in all parts of the genome. Ultraconserved is usually taken to mean that that part of the genome is important (i.e. mutations that happened there were selected against, and so lost from the population at large). That’s why open reading frames are more highly conserved than the areas outside the open reading frames, because mutations in the open reading frames would disrupt the protein and be selected against.
I’m wondering if the researchers checked for effects on longer-term events, such as reproduction?
I’m wondering about more subtle mutations than total deletion of these regions. Is it reasonable to hypothesize that small changes in these regions, for example one or two substitutions, or a small indel, would have effects (“gain of function”) that total deletion would not?
I would expect total deletion to be rather unlikely through spontaneous mutation, so natural selection would never “see” that genotype. But a single-base-pair mutation is not so unlikely, and perhaps has phenotypic effects.
I think this hypothesis is testable, but very labour-intensive, unless some knowledge can be gained about the relative importance of each of these 731 sites in uc467.
There is no real prove that the deletion has really no phenotypic effects. Did they place the mice in all situations imaginable? Are the mice psychologically healthy? What happens if you heat-stress them and then wound them? What about their reaction to any stimulus imaginable?
I cannot really believe that a deletion has no visible effect in the long term.
jimv,
Yes, and according to the synopsis at PLoS Biology: “Their offspring showed no appreciable differences in viability or litter size compared to their control (wild-type) littermates and no anomalies in body weight, age, or survival.”
This paper is very similar to one that I commented on here:
http://interrogatingnature.blogspot.com/2007/07/all-highly-conserved-regions-are.html
Three of the authors are the same on both papers as well.
Ryan,
You are a great blogger because of you sharp eye for controversial issues and a sharp tongue for eloquently voicing your doubtful or occasionally outright negative opinions.
These are important traits for a research professor – but I wonder if you also teach? People are intrigued by controversies, but need the catharsis of some meaningful resolution. Some sort of a “take home lesson†– if you will. With no resolution, titillation may result in confusion, frustration, followed by disinterest and finally boredom.
I loved your “Onion Test†– for any who might think has a coherent concept about the function of “junk DNA†to please explain why different sub-species (e.g. of the onion) show huge differences in the amount of “junk DNA†they have.
I took your challenge of the “Onion Test†in another blog where I provided my explanation based on FractoGene (that is, if junk DNA provides the auxiliary information for recursive fractal development of organelles, organisms and organs, governed by the fractal genes/nongenes) – it follows from fractal algorithms that small changes in their parameters may result in very rapid – or extremely slow – convergence, thus needing small or very large amounts of “Junk DNA” (that of course, is “anything but junk”).
Why don’t we formulate an “Elephant Fish Test†for the enigma of ultraconserved and ultraselected “non-coding DNAâ€? On one hand, such ultraselection was outright called a “Mystery†– while the last 3 authors (Venter, Strausberg, Brenner), out of 12 contributors of the “Elephant fish” paper arrived at the conclusion on conserved noncoding elements (CNEs) that “The ancient vertebrate-specific CNEs in the elephant shark and human genomes are likely to play key regulatory roles in vertebrate gene expressionâ€. Anyone who thinks has a coherent concept about the function of whole genome should be able to provide an explanation of non-coding sequences conserved over half a billion years – yet, at the same time some of these sequences, when eliminated, apparently don’t show up in regular functioning?
Here we go. Let’s see if FractoGene not only can provide an at least as good answer to the “Onion Test†than any other I know of (please let me know if I am missing some) – but let’s see if FractoGene can pass the “Elephant Fish Test�
FractoGene holds that protein-coding DNA (formerly called “genesâ€, before ENCODE necessitated an “update of definition†of both “junk” and “gene”) produce the “materials†of the organisms, while non-(directly)-coding DNA holds the information what is the “design†how they should be put together.
As shown in JunkDNA, some “design†is worth “ultraconserving†for survival (e.g. the vertebral column for vertebrates) – while particular proteins might be substituted.
So how about “not noticing the lack of some important design feature�
Since this is just a blog, let’s take an example that any truck-driver will understand. In the “evolution†of cars, some “design-elements†are “ultra-conserved†(wheels, steering, engine) – while “materials” have been substituted extremely widely. (Hardly any wood and even elimination of glass in late model cars, while none of today’s plastics in early cars, etc., etc.)
Can it happen that one will not detect in the functioning of the automobile if an important “design element” that has been “ultraconserved” is missing?
Of course, it can happen.
For instance, having a spare tire is an “ultraconserved†element of the automobile design through the ages. (As for “survival of the fittest”, who would pick from two comparable models a car that does *not* have a spare tire??)
In normal use (on clean freeways), however, you will not note its absence sometimes for decades.
Taking your car “to the wild” (to unpaved rocky roads, for instance) – is a different matter.
Let’s see if in the scope of a blog, very intelligent “Onion†and “Elephant Fish†tests can be posted – and see if we are getting any “catharsis” (at the least, some attempts at resolutions).
Pellionisz_at_junkdna.com
Andras — instead, why don’t you prepare a manuscript describing in detail the “Fractogene” approach and its predictions and submit it to a journal for peer review. You can request me as a reviewer if you like. Until then, you will probably have trouble winning converts. Blog comment threads certainly are not the place where real science is conducted.
The statement that “these ultraconserved elements make up a small fraction of the total genome of organisms like mammals, on the order of 5%” is incorrect by three orders of magnitude. Ultraconserved elements span several hundred kbp, or about 1% of 1% of a mammalian genome.
“[T]hese regions are known to be located in close proximity to genes for which loss of function mutations result in severe abnormalities.”
Is there any reason at all to suppose that this proximity may have something to do with the degree to which these non-coding regions are conserved?
I have tried to correct my error that has caused some confusion (namely by saying that ultraconserved sequences in particular represent 5% of the genome when this is actually the number for all apparently constrained sequences).
Thank you to aDNAn Derti and Gill Bejerano for pointing this out. I had to fix it on my BlackBerry because I am on the road, so hopefully nothing got messed up.
Ryan,
I already put out several peer-reviewed publications (and was peer-invited to a number of keynote-presentations) on FractoGene. Most relevant to both the “Onion test†and the “Elephant shark test†is the quantitative prediction by FractoGene for the Fugu Purkinje neuron, with experimental results supportive enough such that Dr. Malcolm J. Simons was the lead-author in our publication: Simons, M. and Pellionisz, A. (2006) The Cerebellum.
I also totally agree that blogs are not the place to do real science.
In my view, blogs are truly important (since effective) PR means of communicating real science to „truck drivers†who on one hand wonder where the hell their $30 billion yearly tax dollars go (just for NIH…) – and on the other hand why is it that „the bottom of the Genomics barrel fell outâ€. (In the language of real science: “How to build a new solid set of fundamental concepts?”)
Most importantly, if „truck drivers†(and their families and loved ones) include people with „junk DNA diseasesâ€, we certainly don’t want to project the idea to them that real scientists are a bounch of clawns who ignored 98.7% of the human DNA as „junk†and half of top scientists think that “ultraconserved elements” do nothing, while the other half think that “they serve key roles in regulation”.
We are running the risk of taxpayers instructing their representatives „no more money till these guys fight it out which way is up – I want to stop hunting for nonexistent (and undefined) genes for diseases with their origin most likely in the 98.7%”
While worrying on the appearence of the „scientific establishment†we might also want to stay clear from „number’s gameâ€; if it is X or Y % that is „junkâ€, or „ultraconserved element†in the DNA. Science deserves concepts clarified first.
I am hoping that the „truck driver†will never forget the *concept* that some elements of the design of his car are “ultraconserved” (yet disposable without ill-effects on normal function); whenever he/she looks at – or even thinks of – the spare tire lurking in the trunk …
Pellionisz_at_junkdna.com
Andras, could you please provide a link or a full citation (journal, volume, pages)? That link goes to blogger.com.
I’m interested in reading a more formalized account of the FractoGene hypothesis.
Thanks.
Excellent post! I should have seen this before writing up something on ‘junk DNA’ on my own blog.
The results of the PLoS paper are also very interesting. I am a proponent of the idea that most of the gene-less region is of no use for the individual. Nevertheless, I would have expected that something “ultraconserved” would have some function.
There might be an interesting parallel in the protein world (I am more of a protein person). If you look at sequence similarity of various proteins in two given species (say human and fish), you will see a wide range of conservation. Some are so divergent that similarity is barely visible, some are almost 100% identical. I have tried very hard to find a rationale for having some extremely well conserved genes. However, apart from a few examples there doesn’t seem to be a rule. Many things have been tested, including centrality in interaction networks, but I have never seen a convincing correlation that would explain why the ultraconserved protein sequences are so invariant.
Dear TheBrummell
here is the link and full reference of the peer-reviewed science journal article with experimental support of the predictions of FractoGene (full text can be Googled with ease):
Simons, M, Pellionisz, A; Genomics, morphogenesis and piophysics: Triangulation of Purkinje cell development. The Cerebellum, 2006; 5(1): 27-35
On a related note, I commented upon the importance of *concepts* able to pass the “Onion Test” as well as the “Elephant Shark Test”, and even ventured a remark that NIH support would wane if they don’t move quickly.
Well, they did. The latest entry in junk DNA portal reports that NIH announced today a $210 M program “to develop new ways to think about challenging biomedical problems” in a trans-disciplinary manner… While the re-organization of NIH might take awhile, there are already specific “Junk DNA diseases” targeted.
Now we only need a bit of “theoretical genomics” to put this little “decoding problem” at hand on some new axiomatic grounds… Malcolm Simons said it might only take 500 years…
pellionisz_at_junkdna.com
Andras,
Thanks very much. Weekend reading, ahoy!
Andras —
I looked at the paper. A few comments.
1) This doesn’t really spell out the “FractoGene” approach in general terms, it is more a particular application of it. I still think you need to write a very clear general review of what it is that you are claiming and several clear predictions that it makes.
2) The paper seems to postulate a correlation, though not necessarily a simple linear one, between genome size and the complexity of Purkinje cells. You compare human, guinea pig, zebrafish, and pufferfish. I have commented before about the problems of using only species from the sequencing projects because they are chosen for having small genomes in most cases. You don’t have a frog on there or a salamander, but your prediction would seem to be that P-cell complexity would be much higher in a salamander than in a human. I have not seen the data, but I doubt this will be borne out.
3) You note that the guinea pig P-cell is much less complex than that of human, presumably because it lacks the non-coding “fractal” DNA that humans have. You say in the legend of Fig. 4 “The genome size in the guinea pig is not known to date”. At the time your paper was published, six different estimates for the guinea pig genome size had been published and made available in the database. The guinea pig genome is larger than that of humans. So there is no correlation even within the selective comparison you made.
I have not read the paper in detail, but even on the face of it I would say that the hypothesis is not well supported. Again, blogs are not the place to make your case. You need to 1) spell out the hypothesis unambiguously, and 2) produce data.
Just a brief note: Kay at Suicyte is actually a ‘he’ (look at the beard!). In our parts, most Kays happen to be male. I know that this can be a source of confusion, at least since somebody I once visited in the US suggested to go to a lesbian place when I mentioned that I would be accompanied by my girl-friend.
Ryan,
I am a bit confused. You kindly suggested that I name you as a referee as the FractoGene approach evolves also in publications – yet you say “I looked at the paper … I have not read the paper in detailâ€. I am convinced that you are intrigued by unresolved issues – but less sure if you commit the necessary effort to follow-up (C-value; a general trend along the emergence of species as well as very notable exceptions; onion, salamandra, ultraconserved and ultraselected elements yet no apparent phenotypic aberration in their absence, etc). You could perhaps assign the task to some students (particularly if there are “cross-dominant” types; well-versed both in biology and mathematics) before rushing to judgment. This is important since the FractoGene approach is fairly technical and e.g. in another blog where after rather lengthy debate if there was a better explanation for both the C-value trend *as well to mechanisms of exception* (1) there was no alternative offered as a better approach (2) the bona fide scientist’ last posting (before his closing remarks) revealed a fundamental ill-understanding of fractals (Dr. A.B. confused “fractal dimension” with “fractal complexity” – a very basic error, as mathematically one is a constant the other is a function). Likewise, when you comment on my Fugu paper “This doesn’t really spell out the “FractoGene” approach in general terms, it is more a particular application of it†– I am not sure if you bothered to look up the Fractal model of Purkinje cell paper from which the general power of the fractal approach can be well assessed.
Also, I am puzzled by the comment that “At the time your paper was published, six different estimates for the guinea pig genome size had been published and made available in the database. The guinea pig genome is larger than that of humans.†It is not the date of publication, but date of acceptance that is relevant, and the Apr. 2007 (!) release reveals 1,949,659,178 base-pairs for the cavia porcellus guinea pig. My paper is correct.
Surely, blogs are not relevant for conducting serious science. Suffice to conclude that one still can not see an alternative explanation for the general C-value trend as well as reasons for very notable exception, and for the ultraconserved element “mystery†– other than the fractal approach. Science (if not limited to “data only – please don’t bother me with explanationâ€) does not favor a vacuum in theory. Thus, if the only reasonable (never perfect) explanation is not replaced by a better set of explanation, biology (in this case genomics) remains a “theory-week science”. This is particularly worrisome when Genomics (as we knew it) has apparently lost practically all “axiomatic†(truly, dogmatic) pillars. Those who are really interested, and know better theoretical explanation than what FractoGene approach offers as it evolves, please let me know.
pellionisz_at_junkdna
And for goodness sake, could someone please go read Chris Harrison’s earlier post on Interrogating Nature so he can’t stop the crank-like self promotion in all the discussion threads?”
Ahahaha.
Andras,
You are referring to a rather fragmentary preliminary assembly of the Cavia genome, not to estimates of actual genome size. However, if it is your prediction that the Cavia genome will be around 2Gb according to the expectations of FractoGene, then this is good because it is testable.
It is my opinion that you need to present a clear exposition of the “FractoGene” approach and its predictions, and that this needs to be subjected to peer review and backed up with far more data. I said you could suggest me as a reviewer for this new paper if you wanted. In other words, I am willing to consider the idea if it is presented properly. That is not to say that I think there is much merit to it based on what I have seen, only that I am keeping an open mind. However, if you decline to subject the approach to proper peer review and to provide sufficient data, then I will have little choice but to reject the FractoGene approach along with many other ideas about “function” for non-coding DNA.
The Purkinje cell paper provides a very minimal discussion of the approach. If it is “highly technical”, then you need to make it very clear so that others can evaluate what it states and predicts, and you need a lot of data to support it. I don’t think even the small dataset you selected supports what you claim, but again please clarify for me precisely what genome size you would predict for Cavia and several other groups that have not been measured yet. This is how you test a hypothesis properly.
In short, until you are willing to present the model clearly in a peer-reviewed paper, to make explicit and testable predictions, and to provide data, I am not interested in discussing the “FractoGene” approach any further and I would appreciate it if you would refrain from repeating the same claims over and over on my discussion forums.
Ryan,
“if it is your prediction that the Cavia genome will be around 2Gb according to the expectations of FractoGene, then this is good because it is testable”
Yepp, the “Fugu cerebellum” paper presented a suitable platform (Purkinje neuron), and an approach with an algorithmic handle (fractals) to provide predictions both for the general trend of increasing fractal complexity (and cell size) with genome size, where the algorithmic explanation (fractal growth) allows for the observed huge deviations (rapidly or slowly converging iterative process). For the Fugu Purkinje cell the genome size (1/8 of human) predicted a rudimentary P-cell arborization compared to the human. In case of the two interim species (guinea pig and zebra fish) in one case the P-cell was known but the genome wasn’t, for the zebra fish, the other way around. Now with these quantitative data becoming available neither peer-review publication nor testability are questionable. (On another blog I outlined an experimental protocol way beyond ordinary means of a traditional “not-transdisciplinary” University Department – but more suitable for a Center of PostGenetics.
My blog interference only wished to point out that (a) those who want to know a viable approach were informed about availability (the fractal approach is also helpful to resolve the “mystery” of ultraconserved elements)and (b) help to show “non-obviousness”, (c) discourage people to later say “oh, sure, but it is nothing new – we all knew that all along”.
To show that all is not only about “FractoGene”, I finally mention that there is a rapidly escalating number of scientists who have stepped over the “flabbergasted” stage some time ago and are working feverishly on algorithmic approaches (see under “PostGenetics”). For those considering deep math a friend, I highly recommend Michael F. Barnsley’s “SuperFractals” (2006).
pellionisz_at_junkdna.com
Keeping the response as short as possible, could you confirm that the Purkinje cell paper, and thus the FractoGene approach, assumes that the guinea pig genome size will be roughly intermediate between that of zebrafish and of human?
YES / NO
Specifically, does the FractoGene fail a test if guinea pig total genome size is NOT roughly 2Gb?
YES / NO
I just want to make sure I am clear on what your approach assumes/requires.
Ryan,
No and No. (In your required regretfully short format)
—
P.S.
“could you confirm that the Purkinje cell paper, and thus the FractoGene approach, assumes that the guinea pig genome size will be roughly intermediate between that of zebrafish and of human?”
To this imprecise question the answer should be a precise “Noâ€. (No, I can not confirm an ill-formed statement poised as a question). First; from the question it is unclear if it refers to “Purkinje cell paper 1989” [by Pellionisz] or “Purkinje cell paper 2006†[by Simons and Pellionisz]. Since the 1989 paper does not mention the zebrafish (both papers mention guinea pig and human), by inference one would think, your question refers to the 2006 paper. However, as any reader can tell, the FractoGene approach originated in 2002 and its prediction for fugu (not an assumption for guinea pig) originated in 2003, results started to trickle in 2004 about fugu, paper written in 2005 and peer-reviewed and published in 2006. Your question mixes up both the logic of timeline, and the targeted platforms).
“Specifically, does the FractoGene fail a test if guinea pig total genome size is NOT roughly 2Gb?â€
To this imprecise question the answer should be a precise “Noâ€.
First, the “Purkinje cell paper 2006†is not the FractoGene approach (see above timeline) but one prediction formulated based on the approach (for the arborization of Fugu PC) – and experimentation has proven supportive of the “fugu predictionâ€. Second, the peer-reviewed 2006 paper does not “assume†a “roughly 2Gb†for the guinea pig PC genome. In addition to the experimental support of the fugu PC prediction, the paper left question-marks on Fig. 4. both for the Danio (dendritic arborization) and Guinea Pig (genome sequencing result, reported in September, 2007 as “6x coverage ongoingâ€). It is very usual in the advancement of science that by answering a question further question-marks are raised. While on one hand I understand that sequencing is (still) expensive and takes time, what boggles my mind is that the zebrafish Purkinje arborization could have long been mapped and published (in its most primitive form a student can do the Golgi prep in a week).
Sometimes I wonder if scientists really want solution, or feel that “it is better business†as long as there is none. (Such that they can keep doing “experimentation as usual†without the disruption that paradigm-shifts typically invoke. Since I am quite familiar with this social phenomenon, the first prediction of FractoGene was deliberately composed *not* to require anything but the most pedestrian, routine experimentation). I love this blog, since it points out several controversies of the “establishment” – and this may be one, indeed. (So is the other that in the USA the best interest of medicine is not to cure people – because they would lose a “customer†– but it is better business to get a sick person hooked on some very expensive medicine that has to be taken for the rest of his/her life).
P.P.S.
If there are better explanation(s) for the vexing phenomena floated in this blog than FractoGene can provide, let’s hear them. Those who think FractoGene has predictive power for the fugu PC arborization only, see “Methylation prediction of FractoGene†(not limited to Purkinje cells).
P.P.P.S.
All through my professional life I collaborated with experimentalists both “at arms length” as well as “elbow to elbow”; resulting in a sizable number of peer-reviewed publications . I am presently doing so (of course, we don’t conduct competitive science on blogs), and my openness to collective efforts has been widely expressed (also in Society format), and reiterated herein. The array of challenges has been enormous and “post-Encode†is blown to staggering and humbling. Maybe we should not overly punish those who think outside the box – since suddenly we all are outside the box. Worse, any of us may be diagnosed of some “Junk DNA diseases” of which some I know are actually dying. This explains my more acute interest these days in “fractal defects” in regulatory DNA than the zebrafish PC arbor (that I have seen already, anyway).
pellionisz_at_junkdna.com
Andras,
So much for a short reply. When someone challenges you that the FractoGene approach has not been stated clearly in a peer-reviewed publication, you cite the Purkinje cell paper. When someone challenges the Purkinje cell paper, you say that it is not an exposition of the FractoGene approach.
Frankly, the rhetoric and obfuscation is getting exhausting.
I (and presumably my readers) would appreciate it if you didn’t repeat anything more about the FractoGene approach on this blog.
Ryan,
Sure; I am a guest (only because we shared curiosity about explanation(s) on ultraconserved elements), and will of course honor your request.
Indeed, I intended “blog interference closed” with my “finally” answer before – but found direct questions to me afterwards and thought it would be both impolite and easy to misinterpret if I did not adequately answer. (Readers could stop after “No and No”).
The following might be of interest (just in, a reply to my inquiry to BI about the cavia porcellus projected genome size):
From: “Nicole Davis via RT” questions_at_broad.mit.edu
Date: Wed, 12 Sep 2007 09:19:22 -0400 (EDT)
Dear Dr. Pellionisz,
It’s a bit too early in the project to tell. From a snapshot of one of the last steps of the genome assembly, it looks like it is about 2.7Gb in size – but this could be artificially inflated because of high polymorphism rates in certain regions. The scientists who are working on this should know more definitively in a few weeks.
Hope this information is helpful to you.
Best wishes,
Nicole
Andras,
I was not aware that you had done this. I had tried to give you a fair chance to present your views in a proper scientific discussion, but the fact that you would resort to this makes me wish I hadn’t wasted my time on you.
Ryan,
Self-Defense and Freedom of Expression are often cited as supreme principles of advanced societies. I plead guilty to particular sensitivity to such issues, having lived through the butchered Revolution of 1956 in communist Hungary. (My sincerest gratitude to generous alternatives of oppression offered to my fellow-countrymen and women both by the USA and Canada – in retrospect I wish I had not been stuck in an ultracensored and ultrascontraselected confinement so long, till 1973 when I came as a postdoc to Stanford).
I am also grateful and happy for you that in your blog I have not experienced the “ideological bias and antagonism” that I do believe not only resulted in a quite unbelievable suppression of opinions (let alone denial of funds), but was contrary to the advancement of science beyond the “35 years of Junk DNA imprisonmentâ€. I wrote this up to an “Obituary of Junk DNAâ€, but even that paper was rejected without review, similar to the one co-authored by about 30 IPGS Founders. (Nonetheless, that hastened the release of ENCODE Report from the planned September 2007 to June 14 2007).
My opinion is that my piece about “Malcolm’s story†(“Triumph and Tragedyâ€, after 5 years still never aired in the USA – I don’t know about Canadian release, if any) that in either “antagonistic blog†could have been easily published, due to my skills of fighting oppression, finally got published in both. (Quite a feat, in itself…)
Dear Malcolm (my lead-author in our paper based on Darwinian notion, but the F-word I don’t mention as promised), who has not been given any credit whatsoever (that I know of) in the cacophony of Post-ENCODE-Press and related blogs, may deserve at least as little while his tragically triumphant lifetime lasts.
pellionisz_at_junkdna.com
Found some cool support for my claim (idea is actually from my boss) that mice that have one ultraconserved element deleted and are unchanged doesn’t prove anything: Have a look on this post on PZ Meyer’s blog: If you delete one HOX gene (we’re talking about basic master-genes of development here, right, not some non-coding conserved regions that someone saw in 2003 by looking on the UCSC whole-genome alignments) it doesn’t have any visible effect either. They were all duplicated during evolution and are robust enough of a system.
Read the article above, subsitute “HOX” by “ultraconserved” and you see why I think that we will see phenotypical differences one day, if the right combination of ultras is knocked out.
Here’s how I concluded, Max:
“At most, genomic analysis is showing genome form, function, and evolution to be much too complex to support any inflexible assumptions on either side.”
You can certainly believe whatever you like, but I remain agnostic pending further data.