Leaving aside the difficulty in defining terms such as “complexity” and “gene“, there has been for many decades an underlying assumption that there ought to be some relationship between morphological complexity and the number of protein-coding genes within a genome. This is a holdover from the pre-molecular era of genetics, when it was at first thought that total genome size should be related to gene number, and thus to complexity. Indeed, the constancy of DNA content within chromosome sets (“C-values”) was taken as evidence that DNA is the substance of heredity, and yet it was recognized as early as 1951 that there is no clear relationship between the amount of DNA per genome and organismal complexity (e.g., Mirsky and Ris 1951; Gregory 2005). By 1971, this had become known as the “C-value paradox” because it seemed so self-contradictory (Thomas 1971). (The solution to the C-value paradox was that most eukaryotic DNA is non-coding, although this raises plenty of questions of its own).
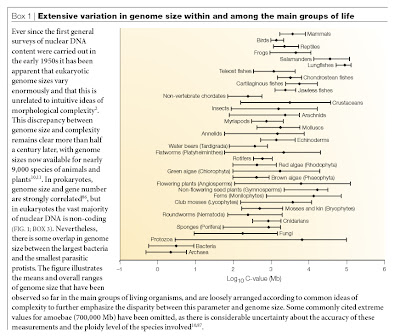
Nevertheless, one sometimes encounters arguments that there is a positive correlation between complexity and genome size, even in the scientific literature. Let me put to rest the notion that genome size is related to complexity on the broad scale of eukaryotic diversity. Here is a figure from Gregory (2005) showing the known ranges and means for more than 10,000 species of animals, plants, fungi, protists, bacteria, and archaea (click image for larger view).
The notion that gene number and complexity should be related has survived largely intact into the post-genomic era, in no small part due to the popular tendency to describe genomes as “blueprints”. Genomes are not blueprints because there is no direct correspondence between a given bit of the genome and a particular piece of the organism. If one must have an analogy for how genomes operate, then a far more appropriate one is with recipes and cakes. No single word in a recipe specifies a particular crumb of a cake, but following the recipe correctly will result in a cake nonetheless. It probably does not need spelling out, but genomes are the recipe, development is the process of mixing ingredients and baking, and organisms are the cake.
Now, one might expect that a more complex cake would require a more verbose recipe, and indeed on a very general level this is true: viruses have very few genes, bacteria and archaea have more, and eukaryotes have more still. Beyond that, however, it is not necessarily the case that a complex cake needs a recipe with more individual instructions. If the language is very efficient — for example, if one sentence in the recipe can convey several steps, or if one can combine the same basic instructions in different ways to make different parts of the cake — then a short recipe might easily produce a more complex cake than one that goes on for several pages.
While predictions regarding human gene number varied considerably prior to the completion of the human genome sequence in 2001, it was nevertheless somewhat surprising that the gene count is only about 20,000-25,000 for a human (International Human Genome Sequencing Consortium 2004). In fact, some people started calling this the “G-value paradox” or “N-value paradox” (for Gene or Number) in reference to the older C-value paradox (Claverie 2001; Betrán and Long 2002; Hahn and Wray 2002).
Here is how Comings (1972) described the C-value paradox:
Being a little chauvinistic toward our own species, we like to think that man is surely one of the most complicated species on earth and thus needs just about the maximum number of genes. However, the lowly liverwort has 18 times as much DNA as we, and the slimy, dull salamander known as Amphiuma has 26 times our complement of DNA. To further add to the insult, the unicellular Euglena has almost as much DNA as man.
And here are Harrison et al. (2002) (probably mostly facetiously):
The sequencing of the genomes of six eukaryotes has provided us with a related quandary: namely, how is the number of genes related to the biological complexity of an organism (termed an ‘N-value’ paradox by Claverie [2001])? How can our own supremely sophisticated species be governed by just 50-100% more genes than the nematode worm?
Of course, neither the “C-value paradox” nor the “G-value paradox” is a paradox at all. As I have said elsewhere, this simply follows the common but erroneous equation of simplistic expectation + contradictory data = “paradoxâ€. Some genes may encode multiple proteins and gene regulation may be more important than gene number, which means that constructing a complex organism does not require a large number of genes any more than it requires a large genome. No paradoxes.
But why might less complex organisms possess large numbers of genes? Rice (Oryza sativa), for example, is thought to have about 50,000 genes, or twice as many as humans (Goff et al. 2002; Yu et al. 2002). One possible explanation is that rice is an ancient polyploid whose entire genome was duplicated in its ancestry. (At least one round of genome duplication also happened early in the evolution of vertebrates, though most lineages now behave genetically as diploids).
But what about something like a purple sea urchin (Strongylocentrotus purpuratus), whose genome apparently encodes 23,300 genes? As deuterostomes, sea urchins are more closely related to vertebrates than to other invertebrates, but that alone does not explain the fact that they have a gene number roughly equivalent to humans (at least, not under the simplified view of genome evolution being discussed). Further, relatedness to self-described complex organisms certainly can’t explain why corals, which are very distant relatives of vertebrates and considered to be relatively “simple” animals, also have somewhere around 20,000 to 25,000 genes.

It turns out that genes involved in immunity are extraordinarily abundant in sea urchins and corals, and that this could account for a significant portion of their total gene number. (Sensory and developmental genes also appear to be very well represented in the sea urchin genome). It is well known that pathogen populations can evolve rapidly and thus that a single host defense mechanism may not remain effective for long. Vertebrates handle the infectious onslaught with a two-tiered system. First, “innate immunity“, which is based on non-specific immune reactions to pathogen attack and is the first response of the body’s immune system. This sort of immunity involves a suite of genes that generate a generalized but limited immune response. In this case there is something of a link with complexity, namely that in order to have a more complex set of possible responses, one would need to have more such genes. All animals possess innate immunity, but only the jawed vertebrates also exhibit “adaptive immunity“, which provides a tailored response to individual pathogens. This system does not involve an individual gene for every possible pathogen, but rather employs an array of duplicated genes that can be shuffled in an effectively limitless number of combinations, like railway cars on a long train, to produce a wide variety combinations of antibodies.
The net result is that vertebrate immunity is more flexible, but that this is achieved not through the addition of tens of thousands of new genes, but through the evolution of a system that can recombine existing genes. Groups like echinoderms and cnidarians, by contrast, may require more immune gene s to accomplish an effective level of defense because they lac
s to accomplish an effective level of defense because they lac k this ability to use existing genes in a large number of combinations. While analogies between human inventions and biological systems can be very problematic, it does seem apt to point out that more sophisticated technologies are frequently simpler, smaller, and more efficient, with fewer parts. A large number of components and a high degree of physical complexity can represent the primitive rather than the derived state in both engineering and evolution.
k this ability to use existing genes in a large number of combinations. While analogies between human inventions and biological systems can be very problematic, it does seem apt to point out that more sophisticated technologies are frequently simpler, smaller, and more efficient, with fewer parts. A large number of components and a high degree of physical complexity can represent the primitive rather than the derived state in both engineering and evolution.
More DNA generally, or more genes in particular, need not relate to morphological complexity. The more knowledge has accumulated about the size, content, and regulation of genomes, the more the basis for expecting such an association has eroded. Being shocked by, or even ashamed of, the fact that humans do not reign supreme in terms of genome size or number of genes is not the appropriate reaction. Rather, realizations such as these should be exciting and should stimulate the next generation of genomic investigation.
_________
References
Betrán, E., and M. Long. 2002. Expansion of genome coding regions by acquisition of new genes. Genetica 115: 65–80.
Claverie, J.-M. 2001. What if there are only 30,000 human genes? Science 291: 1255–1257.
Comings, D.E. 1972. The structure and function of chromatin. Advances in Human Genetics 3: 237-431.
Goff, S.A. et al. 2002. A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296: 92-100.
Gregory, T.R. 2005. Synergy between sequence and size in large-scale genomics. Nature Reviews Genetics 6: 699-708.
Gregory, T.R. 2006. Genomic puzzles old and new. ActionBioScience.org.
Hahn, M.W. and G.A. Wray. 2002. The g-value paradox. Evolution & Development 4: 73-75.
Harrison, P.M., A. Kumar, N. Lang, M. Snyder, and M. Gerstein. 2002. A question of size: the eukaryotic proteome and the problems in defining it. Nucleic Acids Research 30: 1083-1090.
International Human Genome Sequencing Consortium. 2004. Finishing the euchromatic sequence of the human genome. Nature 431: 931–945.
Mirsky, A.E. and H. Ris. 1951. The desoxyribonucleic acid content of animal cells and its evolutionary significance. Journal of General Physiology 34: 451-462.
Pennisi, E. 2006. Sea urchin genome confirms kinship to humans and other vertebrates. Science 314: 908-909.
Rast. J.P., L.C. Smith, M. Loza-Coll, T. Hibino, and G.W. Litman. 2006. Genomic insights into the immune system of the sea urchin. Science 314: 952-956.
Sea Urchin Genome Sequencing Consortium. 2006. The genome of the sea urchin Strongylocentrotus purpuratus. Science 314: 941-952.
Thomas, C.A. 1971. The genetic organization of chromosomes. Annual Review of Genetics 5: 237-256.
Yu et al. 2002. A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296: 92-100.

Our preliminary look at the sea urchin genome suggests that proper annotation is just beginning. My particular genes (HSP70 family) are not in very good shape.
I suggest that the number of genes in the sea urchin genome will drop substantially one the data is cleaned up. That seems to be the trend. The gene finding programs seem to produce over-estimates.
I’d be willing to bet that the final number will be under 20,000.
Thanks for linking to my posting on gene number. We need to get the word out that lots of experts were not surprised when humans were discovered to have only 30,000 genes (or less). It’s the ones who were surprised who are most likely to come up with excuses like alternative splicing and functional RNAs. They seem to have trouble with the concept that humans aren’t far more complex than the “lower” animals that we are supposed to be “superior” to.
Quite right, urchin gene number assessments may shrink somewhat. The larger issue that gene numbers need not be (and generally are not) related to organism complexity would remain, however. I think many people were surprised that the human gene number is *so* low, but to start calling this a paradox is just a repetition of the fallacious reasoning that dogged discussions of genome size several decades earlier.
You’re doing an incredible job! I’m really amazed by bloggers whose posts are full of references. I must add you to my blogroll and feedreader.
Thanks for the kind message. Including references is just part of how one should write science, so I don’t think should be special to see references in blogs that wish to be taken seriously as scientific writing. Maybe that is another part of the experiment for this blog — to see if people will respond well to referenced science writing, or if they only like blogs that link to other blogs or webpages, or don’t include any references whatsoever. We shall see, anyway. 🙂
Then there’s the whole issue of how to measure “complexity”, which Larry hinted at in his comment.
Yep.