In 2001, when the draft sequences were announced, it was revealed that the human genome contains somewhere between 30,000 and 35,000 protein-coding genes (International Human Genome Sequencing Consortium 2001; Venter et al. 2001). The completed sequence, published in 2004, provided an even lower estimate of 20,000 to 25,000 genes (International Human Genome Sequencing Consortium 2004). At present, Ensembl gives the number of protein-coding genes in the human genome as 21,724 known genes plus 1,017 novel genes. (“Known genes” correspond to an identifiable protein, “novel genes” look like they probably correspond to a protein but not yet a known one).
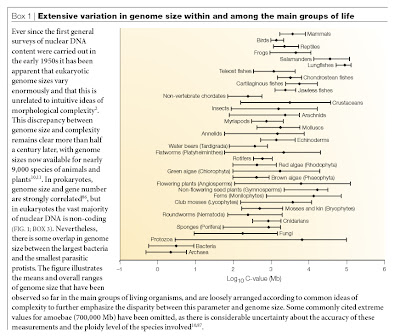
As I have discussed in a previous post, there is quite a bit of interest in comparisons of gene number among species. Part of the reason is that there has long been an expectation that gene number (and prior to the 1970s, genome size) should be linked to some measure of organismal complexity. More often than not, complexity is defined in such a way as to place humans at the top of the scale, but objective metrics also have been attempted. (See the excellent post entitled “Step away from that ladder” by PZ Myers for discussion on this).
Prior to the human genome sequence, the expected gene number most commonly cited was 100,000, even though lower estimates were becoming increasingly common (e.g., Aparicio 2000) and the basis of this figure was somewhat dubious to begin with. As a result, the finding of 20,000-25,000 genes in the human genome has inspired extensive commentary. Some authors even characterized this as a new “G-value paradox” or “N-value paradox”, in reference to the “C-value paradox” of yesteryear (Claverie 2001; Betrán and Long 2002; Hahn and Wray 2002).
Two questions are relevant to this topic: Is the “low” number of protein-coding genes really surprising? If so, is this “paradoxical”?
Between 2000 and 2003, a light-hearted betting pool known as “GeneSweep” was run in which genome researchers could guess at the number of genes in the human genome. A bet placed in 2000 cost $1, but this rose to $5 in 2001 and $20 in 2002 as information about the human genome sequence increased. One had to physically enter the bet in a ledger at Cold Spring Harbor, and all told 165 bets were registered. Bets ranged from 25,497 to 153,438 genes, with a mean of 61,710, as indicated by the plot below.
It has been argued that this shows that a substantial percentage of scientists expected a low gene number and were not surprised by the human gene count estimates. I interpret these data differently, for several reasons.
First, this was a betting pool, and as a result there would have been additional factors influencing the entries. For example, in a sports pool, people may assume that everyone will pick the top-ranked teams and therefore intentionally select an underdog that they hope, but do not necessarily expect, will win. If the most commonly repeated gene count estimate was 100,000 at the time, then this would be the last bet I would have placed. The decision would therefore be to either go higher or lower than this. Personally, I probably would have gone lower rather than higher, because more than 100,000 genes might be problematic due to mutational load. So, based purely on the dynamics of informed betting, I would have expected most people to pick a number substantially lower than 100,000 even if they still believed that to be the most likely number.
Second, it is important to consider when the different bets were placed (I am looking into this out of curiosity). It is entirely possible that the high values were picked first, and then lower numbers were mostly chosen later for two key reasons. One, people had to physically enter their bets at Cold Spring Harbor, so they would have seen what others were guessing and could adjust accordingly (see above). Two, new estimates came out around 2000 that put the value well above 100,000, followed by other estimates that were much closer to 40,000. If the betting trends simply tracked these data, then one could not argue that people always expected a low number. Indeed, it may be that few people would have guessed a low number until very shortly before the release of the sequence.
Third, the winning estimates were higher than the probable total by several thousand genes. (The contest ended in a three way tie, with half of the $1,200 in prize money going to Lee Rowen [who bet 25,947 in 2001] and the other half shared by Paul Dear [27,462 in 2000] and Olivier Jaillon [26,500 in 2002]; see Pennisi 2003, 2007). No one guessed too low. In fact, most entries were far above the high end of the initial draft sequence estimates of 35,000, even though betting continued for at least another year. Likewise, no estimates based on molecular data prior to the close of betting gave a value of 23,000 either.
We may also ask what genome sequencers had to say at the time. James Watson, co-discoverer of the double helix structure of DNA and the original director of the Human Genome Project, wrote the following in 2001:
Until we saw the first DNA scripts underlying multicellular existence, it seemed natural that increasing organismal complexity would involve corresponding increases in gene numbers. So, I and virtually all of my scientific peers were surprised last year when the number of genes of the fruit fly, Drosophila melanogaster, was found to be much lower than that of a less complex animal, the roundworm Caenorhabditis elegans (13,500 vs. 18,500). More shocking still was the recent finding that the small mustard plant, Arabadopsis thaliana, contains many thousand more genes (~28,000) than does C. elegans. Now we are jolted again by the conclusion that the number of human genes may not be much more than 30,000. Until a year ago, I anticipated that human existence would require 70,000-100,000 genes.
J. Craig Venter, who led the private initiatives to sequence the fruit fly and human genomes, was quoted by The Observer in 2001 as saying “When we sequenced the first genome of … the fruit fly, we found it had about 13,000 genes, and we all thought, well we are much bigger and more complicated and so we must have a lot more genes. Now we find that we only have about twice what they have. It makes it a bit difficult to explain the human constitution.” In the same piece, Venter is quoted as noting that “Certainly, it shows that there are far fewer genes than anyone imagined.”
The Human Genome Project Information page said the following in 2004: “This lower estimate came as a shock to many scientists because counting genes was viewed as a way of quantifying genetic complexity. With around 30,000, the human gene count would be only one-third greater than that of the simple roundworm C. elegans at about 20,000 genes”.
Lee Rowen, co-winner of GeneSweep, noted that her estimate was inspired by Jean Weisenbach of Genoscope, who had suggested a few years earlier that the human gene number might be low. Rowen noted that, “at the time, everybody nearly fell of their chair” upon hearing this proposition (Pennisi 2003).
The list could go on, but I think it is evident that, in the light of pre-genomic views of genetics, most scientists were surprised by the low gene count in humans, especially when compared to other species to which we intuitively attribute lesser complexity. Notably, the nematode Caenorhabditis elegans appears to possess 20,069 known and novel genes even though it consists only of ~1,000 cells, the fly Drosophila melanogaster has 14,039 genes, the sea urchin Strongylocentrotus purpuratus 23,300 on first pass, and rice Oryza sativa upwards of 50,000. I don’t know if anyone expected this pattern prior to the dawn of genome sequencing, but I personally have not met him or her.
The question, then, is how will this discrepancy between expectation and reality be resolved? Is it truly “paradoxical”, meaning that it is self-contradictory? Or is it simply a matter of updating our understanding of how genetics works? Coming from the field of genome size evolution, which underwent the same transition decades earlier, I am of the view that “paradox” is not an appropriate descriptor. A complex puzzle — a “G-value enigma” — it may be, but it is one that can be resolved with a broader approach to genetics and much additional research. It took several decades for it to become widely acknowledged that genome size and gene number are unrelated (and indeed, a few authors still argue against it based on a biased dataset consisting exclusively of small, sequenced genomes; see Gregory 2005 for discussion), but we are now developing a reasonable understanding of the non-coding elements of the genome that make up the difference, their effects and (sometimes) functions, and their evolutionary dynamics. Similarly, there are many reasons why gene number and complexity need not be correlated. A list of possibilities is available here (though it was compiled for rather different reasons).
The expectation seems to have been that humans should have comparatively high gene numbers. We do not, and at first this was surprising. Now let us move on to a post-genomic understanding of genetics, and focus less on counting one-dimensional parameters and more on appreciating and ultimately deciphering the complexity inherent within the genome.
_____________
References
Anonymous. 2000. The nature of the number. Nature Genetics 25: 127-128.
Aparicio, S.A.J.R. 2000. How to count…human genes. Nature Genetics 25: 129-130.
Betrán, E. and M. Long. 2002. Expansion of genome coding regions by acquisition of new genes. Genetica115: 65-80.
Claverie, J.-M. 2001. What if there are only 30,000 human genes? Science 291: 1255-1257.
Dunham, I. 2000. The gene guessing game. Yeast 17: 218-224.
Gregory, T.R. 2005. Synergy between sequence and size in large-scale genomics. Nature Reviews Genetics 6: 699-708.
Hahn, M.W. and G.A. Wray. 2002. The g-value paradox. Evolution & Development 4: 73-75.
International Human Genome Sequencing Consortium. 2001. Initial sequencing and analysis of the human genome. Nature 409: 860-921.
International Human Genome Sequencing Consortium. 2004. Finishing the euchromatic sequence of the human genome. Nature 431: 931-945.
Pennisi, E. 2000. And the gene number is …? Science 288: 1146-1147.
Pennisi, E. 2003. A low gene number wins the GeneSweep pool. Science 300: 1484.
Pennisi, E. 2007. Working the (gene count) numbers: finally, a firm answer? Science 316: 1113.
Pennisi, E. 2007. Why do humans have so few genes? Science 309: 80.
Semple, C.A.M., K.L. Evans, and D.J. Porteous. 2001. Twin peaks: the draft human genome sequence. Genome Biology 2: comment2003.2001-comment2003.2005.
Venter, J.C., et al. 2001. The sequence of the human genome. Science 291: 1304-1351.
Watson, J.D. 2001. The human genome revealed. Genome Research 11: 1803-1804.