“It seems as though ‘junk DNA’ has become a legitimate jargon in a glossary of molecular biology. Considering the violent reactions this phrase provoked when it was first proposed in 1972, the aura of legitimacy it now enjoys is amusing, indeed.â€
The origin of “junk DNA”
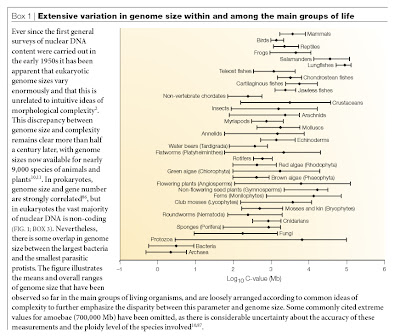
Two main problems struck Susumu Ohno as particularly important in his seminal work on the genetics of evolutionary diversification. The first was the lack of correspondence between genome size (amount of DNA) and morphological complexity (taken as a proxy for gene number), which was a prominent topic of discussion in the early 1970s. As he noted in 1972, “If we take the simplistic assumption that the number of genes contained is proportional to the genome size, we would have to conclude that 3 million or so genes are contained in our genome. The falseness of such an assumption becomes clear when we realize that the genome of the lowly lungfish and salamanders can be 36 times greater than our own†(Ohno 1972a). In fact, Ohno and his colleagues were well aware that much of the DNA in the mammalian genome could not code for proteins, lest the mutational load become fatally high (e.g., Comings 1972; Ohno 1972b, 1974).
The second problem related to the conservative force of purifying selection and the limitations it places on the diversification of species. Ohno (1973) attempted to kill both of these vexatious birds with a single conceptual stone:
The points I wish to make are: 1) Natural selection is an extremely conservative force. So long as a particular function is assigned to a single gene locus in the genome, natural selection only permits trivial mutations of that locus to accompany evolution. 2) Only a redundant copy of a gene can escape from natural selection and while being ignored by natural selection can accumulate meaningful mutation to emerge as a new gene locus with a new function. Thus, evolution has been heavily dependent upon the mechanism of gene duplication. 3) The probability of a redundant copy of an old gene emerging as a new gene, however, is quite small. The more likely fate of a base sequence which is not policed by natural selection is to become degenerate. My estimate is that for every new gene locus created about 10 redundant copies must join the ranks of functionless DNA base sequence. 4) As a consequence, the mammalian genome is loaded with functionless DNA.
Only a redundant copy of a gene can escape from natural selection and while being ignored by natural selection can accumulate meaningful mutation to emerge as a new gene locus with a new function. Thus, evolution has been heavily dependent upon the mechanism of gene duplication. 3) The probability of a redundant copy of an old gene emerging as a new gene, however, is quite small. The more likely fate of a base sequence which is not policed by natural selection is to become degenerate. My estimate is that for every new gene locus created about 10 redundant copies must join the ranks of functionless DNA base sequence. 4) As a consequence, the mammalian genome is loaded with functionless DNA.
The corpulent genomes of dipnoans and urodele amphibians were similarly thus accounted for under this view: “Lungfish and salamanders clearly show the tragic consequences of exclusive dependence upon tandem duplication†(Ohno 1970, p.96). Of course, this differs from current thinking about lungfish and salamander genome size, but that’s another story.
To Ohno, this situation not only permitted, but also paralleled, the evolution of life at large. As he put it, “The earth is strewn with fossil remains of extinct species; is it any wonder that our genome too is filled with the remains of extinct genes?†(Ohno 1972a). The primary outcome of this gene duplication mechanism would not be the generation of new genes, but the deactivation of redundant copies – just as extinction has been the fate of more than 99% of species that have ever lived (Raup 1991). Once purifying selection ceased to shelter gene sequences from change, they would be free to mutate and, if one imagines a set of three gene copies initially sharing the same sequence, it is likely that “in a relatively short time, two of the three duplicates would join the ranks of ‘garbage DNA’†(Ohno 1970, p.62).
In Ohno’s usage, as in the vernacular, “garbage†refers to both the loss of function and the lack of any further utility (it was once useful, but now it isn’t). “Garbage DNA†proved to be an unsuccessful meme, but its essence remains in the wildly popular term coined by Ohno two years later – “junk DNAâ€. Thus, as Ohno (1972b) stated, “at least 90% of our genomic DNA is ‘junk’ or ‘garbage’ of various sortsâ€. Interestingly, Ohno mentioned “junk DNA†only in the titles of two of his papers (1972a, 1973), and invoked the term only once in passing in a third (1972b). Comings (1972), on the other hand, gave what must be considered the first explicit discussion of the nature of “junk DNAâ€, and was the first to apply the term to all non-coding DNA.
There are several independent mechanisms by which non-coding DNA can accumulate in the genome. Gene duplication and deactivation is one such mechanism, but this, we now know, applies to only a minority of the non-coding sequences. Nevertheless, the term “junk DNA†was used in some early general descriptions of non-coding elements, including heterochromatin. For example, Comings (1972) noted that:
It has frequently been suggested that the DNA of genetically inactive heterochromatin represents the degenerate and useless DNA of the genome. However, heterochromatin rarely constitutes more than 20% of the genome. This suggests that there are two categories of junk DNA, (1) DNA of constitutive heterochromatin which is neither transcribed nor translated, and (2) nonheterochromatic junk DNA which is probably transcribed, but not translated. This distinction adds one more dimension to the mystery of heterochromatic DNA. Why is it singled out to be nontranscribable when being nontranslatable seems adequate for most of the junk DNA? Perhaps there is clustered junk (heterochromatic DNA) and nonclustered junk, just as there is clustered repetitious DNA (satellite DNA) and nonclustered repetitious DNA.
Later, Ohno himself began applying the term “junk†to heterochromatic, intergenic, and intronic sequences: “Much of this junk DNA occurs as large heterochromatin blocks, often localized in pericentric regions of mammalian chromosomes, or as intergenic spacers and intervening sequences within genes.†(Ohno 1985).
It is clear, however, that Ohno (1982) believed all these sequences were produced by gene duplication:
This great preponderance of intergenic spacers in the euchromatic region is due mostly to the extreme inefficacy of the mechanism of gene duplication as a means of creating new genes with altered active sites. For every redundant copy of the pre-existent gene that emerged triumphant as a new gene, hundreds of other copies must have degenerated to join the rank of junk DNA.
This mechanism alone was considered capable of explaining the vast intergenic regions of eukaryotic genomes. According to Ohno (1985):
Indeed, the abundance of pseudogenes (recent degenerates) attests to the inefficacy of gene duplication as a means of acquiring new genes with novel functions. The net consequence of hundreds of millions of years of continuous gene duplication is the desertification of the euchromatic region of modern vertebrates; the average distance between still functioning gene loci becoming progressively longer.
Junk DNA, function, and non-function
“Junk DNA†had a specific meaning when it first was formulated. It was meant to describe the loss of protein-coding function by deactivated gene duplicates, which in turn were believed to constitute the bulk of eukaryotic genomes. As different types of non-coding DNA were identified, the concept of gene duplication as their source – and therefore “junk DNA†as their descriptor – found new and broader application. However, it is now clear that most non-coding DNA is not produced by this mechanism, and is therefore not accurately described as “junk†in the original sense.
The term “pseudogene†— the technical term for functionless gene copies — was not coined until 1977 (Jacq et al. 1977), and the more explicit definition of these sequences that specified non-function in terms of protein-coding emerged almost a decade later. So, although Ohno’s original description of “junk DNA†obviously involved what are now called “pseudogenesâ€, there was no initial requirement for non-function. As Comings (1972) put it, “Being junk doesn’t mean it is entirely useless. Common sense suggests that anything that is completely useless would be discarded.†(This is what Sydney Brenner meant by the distinction between “trash” or “rubbish”, which one throws away, and “junk”, which one keeps; Brenner 1998). Of course, Ohno did reject the notion of protein-coding function for the extinct genes. As he described it, “a functional gene locus is defined as that DNA base sequence which may sustain deleterious mutationsâ€, and from this it followed that “a DNA base sequence in which all sorts of mutational changes are permissible is obviously not contributing to the well-being of an organism, and for this very reason, it has no function†(Ohno 1973). On the other hand, and in the same publication, Ohno (1973) suggested a different role for non-coding DNA: “The bulk of functionless DNA in the mammalian genome may serve as a damper to give a reasonably long cell generation time (12 hours or so instead of several minutes)â€.
From the very beginning, the concept of “junk DNA†has implied non-functionality with regards to protein-coding, but left open the question of sequence-independent impacts (perhaps even functions) at the cellular level. “Junk DNA” may now be taken to imply total non-function and is rightly considered problematic for that reason, but no such tacit assumption was present in the term when it was coined.
Two groups of people, though maximally divergent in their reasons for so doing, have been driven by a philosophical need to identify functions for all non-coding DNA. The first includes strict adaptationists, among whom it was often assumed that all non-coding DNA, by virtue of its very existence, must be endowed with some as-yet-unknown function of critical importance: “The very fact that amplified sequences have been maintained, withstanding rigours of selection, indicates some adaptive significance†(Sharma 1985).
We may also consider the following discussion comments recorded at the end of Ohno (1973):
Yunis: “This is what I emphasized earlier, that this DNA must have a functional value since nothing is known so widespread and universal in nature that has proven useless.â€
Fraccaro: “Well, there is an exception to that rule. A lot of us have permanent positions at the University but are considered by others (mainly by students) meaningless and of no utility whatsoever.â€
These examples aside, it seems likely that most evolutionary biologists today could tolerate a conclusion, if such were rendered, that a significant fraction of non-coding DNA is functionless. This is not true of the second group in question, compared to whom the passion for function is unrivaled. As Dawkins (1999) suggested, “creationists might spend some earnest time speculating on why the Creator should bother to litter genomes with untranslated pseudogenes and junk tandem repeat DNAâ€. In fact, many have done so (e.g., Gibson 1994; Wieland 1994; Batten 1998; Jerlström 2000; Walkup 2000; Woodmorappe 2000; Bergman 2001). Although apparently “not enough is yet known about eukaryotic genomes to construct a comprehensive creationist model of pseudogenes†(Woodmorappe 2000), the theme that undergirds all of these discussions is that all non-coding DNA must, a priori, be functional.
To satisfy this expectation, creationist authors (borrowing, of course, from the work of molecular biologists, as they do no such research themselves) simply equivocate the various types of non-coding DNA, and mistakenly suggest that functions discovered for a few examples of some types of non-coding sequences indicate functions for all (see Max 2002 for a cogent rebuttal to these creationist confusions). Case in point: a few years ago, much ado was made of Beaton and Cavalier-Smith’s (1999) titular proclamation, based on a survey of cryptomonad nuclear and nucleomorphic genomes, that “eukaryotic non-coding DNA is functionalâ€. The point was evidently lost that the function proposed by Beaton and Cavalier-Smith (1999) was based entirely on coevolutionary interactions between nucleus size and cell size.
Those who complain about a supposed unilateral neglect of potential functions for non-coding DNA simply have been reading the wrong literature. In fact, quite a lengthy list of proposed functions for non-coding DNA could be compiled (for an early version, see Bostock 1971). Examples include buffering against mutations (e.g., Comings 1972; Patrushev and Minkevich 2006) or retroviruses (e.g., Bremmerman 1987) or fluctuations in intracellular solute concentrations (Vinogradov 1998), serving as binding sites for regulatory molecules (Zuckerkandl 1981), facilitating recombination (e.g., Comings 1972; Gall 1981; Comeron 2001), inhibiting recombination (Zuckerkandl and Hennig 1995), influencing gene expression (Britten and Davidson 1969; Georgiev 1969; Nowak 1994; Zuckerkandl and Hennig 1995; Zuckerkandl 1997), increasing evolutionary flexibility (e.g., Britten and Davidson 1969, 1971; Jain 1980; reviewed critically in Doolittle 1982), maintaining chromosome structure and behaviour (e.g., Walker et al. 1969; Yunis and Yasmineh 1971; Bennett 1982; Zuckerkandl and Hennig 1995), coordingating genome function (Shapiro and von Sternberg 2005), and providing multiple copies of genes to be recruited when needed (Roels 1966).
Does non-coding DNA have a function? Some of it does, to be sure. Some of it is involved in chromosome structure and cell division (e.g., telomeres, centromeres). Some of it is undoubtedly regulatory in nature. Some of it is involved in alternative splicing (Kondrashov et al. 2003). A fair portion of it in various genomes shows signs of being evolutionarily conserved, which may imply function (Bejerano et al. 2004; Andolfatto 2005; Kondrashov 2005; Woolfe et al. 2005; Halligan and Keightley 2006). On the other hand, the largest fraction is comprised of transposable elements — some of which become co-opted by the host genome, some of which play major role in generating genomic variation, some of which may be involved in cellular stress response, and yet others of which remain detrimental to host fitness (Kidwell and Lisch 2001; Biémont and Vieira 2006). The upshot is that some non-coding DNA is most c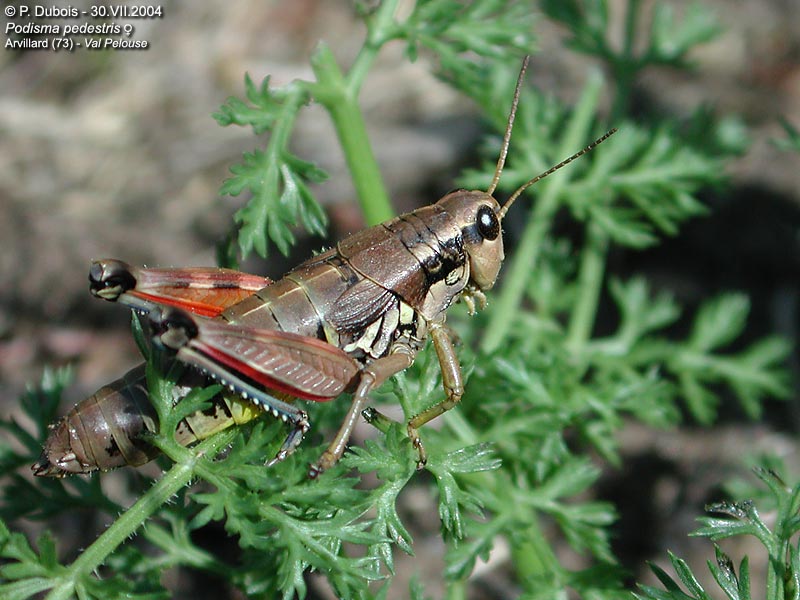 ertainly functional — but when it is, this usually makes sense only in an evolutionary context, particularly through processes like co-option. More broadly, those who would attribute a universal function for non-coding DNA must bear the following in mind: any proposed function for all non-coding DNA must explain why an onion or a grasshopper needs five times more of it than anyone reading this sentence.
ertainly functional — but when it is, this usually makes sense only in an evolutionary context, particularly through processes like co-option. More broadly, those who would attribute a universal function for non-coding DNA must bear the following in mind: any proposed function for all non-coding DNA must explain why an onion or a grasshopper needs five times more of it than anyone reading this sentence.
Should “junk†be thrown out?
There is nothing wrong with a word taking on a new meaning as knowledge changes – that is, unless reference to an original (and outmoded) sense lingers as a source of confusion, or the term expands so much as to lose contact with an initially accurate definition. Indeed, even the term “evolution†is technically a misnomer since its etymology implies an “unfoldingâ€, as of a pre-determined developmental program (see Bowler 1975). The objection raised here is not to terms that change in usage per se, but to those whose shifting usage involves collecting or retaining unwanted conceptual baggage. This is especially relevant when the baggage is toted surreptitiously (note that no serious biologist takes “evolution†to mean a pre-determined unfolding but that ideas of inherent “progress†have been almost impossible to shake; see Gould 1996; Ruse 1996).
“Junk DNAâ€, which originally was coined in reference to now-functionless gene duplicates (i.e., true broken-down “junkâ€), is now used as “a catch-all phrase for chromosomal sequences with no apparent function†(Moore 1996). Its current usage also implies a lack of function which is accurate by definition for pseudogenes in regard to protein-coding, but which does not hold for all non-coding elements. The term has deviated from or outgrown its original use, and its continued invocation is non-neutral in its expression – and generation – of conceptual biases.
“Junk DNA” is not the only offender. Non-coding DNA has been called by many names that have had the same pejorative undertones (intentional or not) implying uselessness, if not outright wastefulness. Examples include excess DNA (Zuckerkandl 1976; Doolittle and Sapienza 1980), surplus or nonessential or degenerate or silent DNA (Comings 1972; Gilbert 1978), quiet DNA (Lefevre 1971), garbage DNA (Ohno 1970), non-informational or nonsense DNA (Ohno 1972b), worthless DNA (Ohno 1973), trivial DNA (Ohno 1974), vestigial DNA (Loomis 1973), redundant DNA (Vinogradov 1998), supplementary DNA (Hutchinson et al. 1980), secondary DNA (Hinegardner 1976), and incidental DNA (Jain 1980).
As Gould (2002, p.503) stated, “A rose may retain its fragrance under all vicissitudes of human taxonomy, but never doubt the power of a name to shape and direct our thoughtsâ€. Because it is generally no longer applied in its original meaningful sense, because the type of DNA to which it actually relates now has a more descriptive name (pseudogenes), and because of its connotations of total phenotypic inertness, the term “junk DNA†should probably be abandoned in favour of less subjective terminology. “Non-coding DNA” serves this purpose quite well.
Concluding remarks
It is an exciting time in genome biology. Aspects of genomic form and function that were largely inconceivable only a few decades ago are now being revealed on a daily basis. It should come as no surprise (and indeed, it probably does not) that new roles are being discovered for non-coding DNA and that some of yesterday’s buzzwords — including “junk DNA” — are destined for the dustbin. However, extrapolating each report that a given small segment of DNA may be functional to mean that all non-coding DNA is vital is as counterproductive as dismissing non-coding DNA as totally non-functional. Genomes are complex, and there is little use in approaching them from a simplistic point of view.
——
Andolfatto, P. 2005. Adaptive evolution of non-coding DNA in Drosophila. Nature 437: 1149-1152.
Batten, D. 1998. ‘Junk’ DNA (again). Creation Ex Nihilo Technical Journal 12: 5.
Beaton, M.J. and T. Cavalier-Smith. 1999. Eukaryotic non-coding DNA is functional: evidence from the differential scaling of cryptomonad genomes. Proceedings of the Royal Society of London, Series B 266: 2053-2059.
Bejerano, G., M. Pheasant, I. Makunin, S. Stephen, W.J. Kent, J.S. Mattick, and D. Haussler. 2004. Ultraconserved elements in the human genome. Science 304: 1321-1325.
Bennett, M.D. 1982. Nucleotypic basis of the spatial ordering of chromosomes in eukaryotes and the implications of the order for genome evolution and phenotypic variation. In Genome Evolution (eds. G.A. Dover and R.B. Flavell), pp. 239-261. Academic Press, New York.
Bergman, J. 2001. The functions of introns: from junk DNA to designed DNA. Perspectives on Science and Christian Faith 53: 170-178.
Biémont, C. and C. Vieira. 2006. Junk DNA as an evolutionary force. Nature 443: 521-524.
Bostock, C. 1971. Repetitious DNA. Advances in Cell Biology 2: 153-223.
Bowler, P.J. 1975. The changing meaning of “evolution”. Journal of the History of Ideas 36: 95-114.
Bremmerman, H.J. 1987. The adaptive significance of sexuality. In The Evolution of Sex and its Consequences (ed. S.C. Stearns), pp. 135-161. Birkhauser Verlag, Basel.
Brenner, S. 1998. Refuge of spandrels. Current Biology 8: R669.
Britten, R.J. and E.H. Davidson. 1969. Gene regulation for higher cells: a theory. Science 165: 349-357.
Britten, R.J. and E.H. Davidson. 1971. Repetitive and non-repetitive DNA sequences and a speculation on the origins of evolutionary novelty. Quarterly Review of Biology 46: 111-138.
Castillo-Davis, C.I. 2005. The evolution of noncoding DNA: how much junk, how much func? Trends in Genetics 21: 533-536.
Comeron, J.M. 2001. What controls the length of noncoding DNA? Current Opinion in Genetics & Development 11: 652-659.
Comings, D.E. 1972. The structure and function of chromatin. Advances in Human Genetics 3: 237-431.
Dawkins, R. 1999. The “information challenge”: how evolution increases information in the genome. Skeptic 7: 64-69.
Doolittle, W.F. and C. Sapienza. 1980. Selfish genes, the phenotype paradigm and genome evolution. Nature 284: 601-603.
Doolittle, W.F. 1982. Selfish DNA after fourteen months. In Genome Evolution (eds. G.A. Dover and R.B. Flavell), pp. 3-28. Academic Press, New York.
Gall, J.G. 1981. Chromosome structure and the C-value paradox. Journal of Cell Biology 91: 3s-14s.
Georgiev, G.P. 1969. On the structural organization of operon and the regulation of RNA synthesis in animal cells. Journal of Theoretical Biology 25: 473-490.
Gibbs, W.W. 2003. The unseen genome: gems among the junk. Scientific American 289(5): 46-53.
Gibson, L.J. 1994. Pseudogenes and origins. Origins 21: 91-108.
Gilbert, W. 1978. Why genes in pieces? Nature 271: 501.
Gould, S.J. 1996. Full House. Harmony Books, New York.
Gould, S.J. 2002. The Structure of Evolutionary Theory. Harvard University Press, Cambridge, MA.
Halligan, D.L. and P.D. Keightley. 2006. Ubiquitous selective constraints in the Drosophila genome revealed by a genome-wide interspecies comparison. Genome Research 16: 875-884.
Hinegardner, R. 1976. Evolution of genome size. In Molecular Evolution (ed. F.J. Ayala), pp. 179-199. Sinauer Associates, Inc., Sunderland.
Hutchinson, J., R.K.J. Narayan, and H. Rees. 1980. Constraints upon the composition of supplementary DNA. Chromosoma 78: 137-145.
Jacq, C., J.R. Miller, and G.G. Brownlee. 1977. A pseudogene structure in 5S DNA of Xenopus laevis. Cell 12: 109-120.
Jain, H.K. 1980. Incidental DNA. Nature 288: 647-648.
Jerlström, P. 2000. Pseudogenes: are they non-functional? Creation Ex Nihilo Technical Journal 14: 15.
Kidwell, M.G. and D.R. Lisch. 2001. Transposable elements, parasitic DNA, and genome evolution. Evolution 55: 1-24.
Kondrashov, F.A. and E.V. Koonin. 2003. Evolution of alternative splicing: deletions, insertions and origin of functional parts of proteins from intron sequences. Trends in Genetics 19: 115-119.
Kondrashov, A.S. 2005. Fruitfly genome is not junk. Nature 437: 1106.
Lefevre, G. 1971. Salivary chromosome bands and the frequency of crossing over in Drosophila melanogaster. Genetics 67: 497-513.
Loomis, W.F. 1973. Vestigial DNA? Developmental Biology 30: F3-F4.
Makalowski, W. 2003. Not junk after all. Science 300: 1246-1247.
Max, E.E. 2002. Plagiarized errors and molecular genetics: another argument in the evolution-creation controversy. Talk.Origins Archive.
Moore, M.J. 1996. When the junk isn’t junk. Nature 379: 402-403.
Nowak, R. 1994. Mining treasures from ‘junk DNA’. Science 263: 608-610.
Ohno, S. 1970a. Evolution by Gene Duplication. Springer-Verlag, New York.
Ohno, S. 1970b. The enormous diversity in genome sizes of fish as a reflection of nature’s extensive experiments with gene duplication. Transactions of the American Fisheries Society 1970: 120-130.
Ohno, S. 1972. So much “junk” DNA in our genome. In Evolution of Genetic Systems (ed. H.H. Smith), pp. 366-370. Gordon and Breach, New York.
Ohno, S. 1973. Evolutional reason for having so much junk DNA. In Modern Aspects of Cytogenetics: Constitutive Heterochromatin in Man (ed. R.A. Pfeiffer), pp. 169-173. F.K. Schattauer Verlag, Stuttgart, Germany.
Ohno, S. 1974. Chordata 1: protochordata, cyclostomata, and pisces. In Animal Cytogenetics, Vol. 4 (ed. B. John), pp. 1-92. Gebrüder Borntraeger, Berlin.
Ohno, S. 1982. The common ancestry of genes and spacers in the euchromatic region: omnis ordinis hereditarium a ordinis priscum minutum. Cytogenetics and Cell Genetics 34: 102-111.
Ohno, S. 1985. Dispensable genes. Trends in Genetics 1: 160-164.
Patrushev, L.I. and I.G. Minkevich. 2006. Eukaryotic noncoding DNA sequences provide genes with an additional protection against chemical mutagens. Russian Journal of Bioorganic Chemistry 32: 1068-1620.
Petsko, G.A. 2003. Funky, not junky. Genome Biology 4: 104.
Raup, D.M. 1991. Exctinction. W.W. Norton & Co., New York.
Roels, H. 1966. “Metabolic” DNA: a cytochemical study. International Review of Cytology 19: 1-34.
Ruse, M. 1996. Monad to Man. Harvard University Press, Cambridge, MA.
Shapiro, J.A. and R. von Sternberg. 2005. Why repetitive DNA is essential to genome function. Biological Reviews 80: 227-250.
Sharma, A.K. 1985. Chromosome architecture and additional elements. In Advances in Chromosome and Cell Genetics (eds. A.K. Sharma and A. Sharma), pp. 285-293. Oxford and IBH Publishing Co., New Delhi.
Slack, F.J. 2006. Regulatory RNAs and the demise of ‘junk’ DNA. Genome Biology 7: 328.
Vinogradov, A.E. 1998. Buffering: a possible passive-homeostasis role for redundant DNA. Journal of Theoretical Biology 193: 197-199.
Walker, P.M.B., W.G. Flamm, and A. McLaren. 1969. Highly repetitive DNA in rodents. In Handbook of Molecular Cytology (ed. A. Lima-de-Faria), pp. 52-66. North-Holland Publishing Co., Amsterdam.
Walkup, L.K. 2000. Junk DNA: evolutionary discards or God’s tools? Creation Ex Nihilo Technical Journal 14: 18-30.
Wickelgren, I. 2003. Spinning junk into gold. Science 300: 1646-1649.
Wieland, C. 1994. Junk moves up in the world. Creation Ex Nihilo Technical Journal 8: 125.
Woodmorappe, J. 2000. Are pseudogenes ‘shared mistakes’ between primate genomes? Creation Ex Nihilo Technical Journal 14: 55-71.
Woolfe, A., M. Goodson, D.K. Goode, P. Snell, G.K. McEwen, T. Vavouri, S.F. Smith, P. North, H. Callaway, K. Kelly, K. Walter, I. Abnizova, W. Gilks, Y.J.K. Edwards, J.E. Cooke, and G. Elgar. 2005. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biology 3: e7.
Yunis, J.J. and W.G. Yasmineh. 1971. Heterochromatin, satellite DNA, and cell function. Science174: 1200-1209.
Zuckerkandl, E. 1976. Gene control in eukaryotes and the C-value paradox: “Excess” DNA as an impediment to transcription of coding sequences. Journal of Molecular Evolution 9: 73-104.
Zuckerkandl, E. and W. Hennig. 1995. Tracking heterochromatin. Chromosoma 104: 75-83.
Zuckerkandl, E. 1997. Junk DNA and sectorial gene expression. Gene 205: 323-343.
__________
Update: At Sandwalk, Larry Moran argues that the term “junk DNA” is “a good term”, “an accurate term”, and “a useful term”. You can read my response in the comments section of the original post or in my re-post on this blog.











Ryan Gregory has serious doubts about the usefulness of the term as he explains in his excellent article A word about “junk DNA”.
Just to clarify, I think the term could be useful — indeed, it was useful when Ohno coined it. The problem is that it is seldom used in an appropriate way. If the meaning were specified explicitly to be “regions strongly suspected of being non-functional with evidence to back it up” (which, incidentally, is not the original definition according to Ohno (1972) or Comings (1972)), and if people used it only in this way, then I would not have a problem with this. But given the difficulty that people seem to have in accepting that some DNA may truly not have a function at the organism level, I don’t know if we could ever get it to be used with such precision.
…a new term, Junctional DNA, to describe DNA that probably has a function but that function isn’t known… think we don’t need to go there. It’s sufficient to remind people that lots of DNA outside of genes has a function and these functions have been known for decades.
That neologism was suggested in response to Minkel’s appeal for a term that would “make the distinction between functional and nonfunctional noncoding DNA clear to a popular audience”. My main suggestion was to call DNA by what it is known to be, if at all possible, by function (“regulatory DNA”, “structural DNA”) or by type (“pseudogene”, “transposable element”, “intron”). Your definition of “junk DNA” is also more precise than most usages, meaning that you specify that the term only be applied to sequences for which there is evidence (not just assumption) of non-function. That leaves us with something in between for journalists to talk about with a catchy buzzword. “Junctional DNA” lets them specify that we’re not talking about “junk DNA” or “functional DNA” — i.e., there is some evidence for function (e.g., being conserved) but no evidence of what that function is. The main utility would be to stop the very frustrating leap that gets made from “this 1% of the genome may have a function, so the whole thing must have this function” kind of reporting. Now they could say “another 1% has moved into the category of ‘junctional DNA'”. I think that would be considerably less misleading than current wording.
Note that I’m avoiding the term “noncoding” DNA here. This is because to me the term “coding DNA” only refers to the coding region of a gene that encodes a protein … there are many genes for RNAs that are not properly called coding regions so they would fall into the noncoding DNA category … introns in eukaryotic genomes would be “noncoding DNA” as far as I’m concerned. I think that Ryan Gregory and others use the term “noncoding DNA” to refer to all DNA that’s not part of a gene instead of all DNA that’s not part of the coding region of a protein encoding gene. I’m not certain of this.
By definition, non-coding DNA is, and always has been, everything other than exons. The reason this is relevant is that early work in genome biology assumed that there should be a 1 to 1 correspondence between DNA content and protein-coding gene number. This is work that occurred for at least two decades before the discovery of introns, pseudogenes, and other non-coding DNA. Now we have more descriptive names for the categories of DNA that are not the genes, all the genes, and nothing but the genes. I actually don’t know of anyone else who would have a problem calling introns, pseudogenes, and regulatory regions “non-coding DNA”. Certainly, Ohno, Crick, and many others have historically put introns in the same non-protein-coding grouping as pseudogenes. It’s just a category — you also have more specific subcategories to apply to each of the types of non-coding DNA. Perhaps your objection relates to an undue emphasis on the distinction between exons and everything else — well, that’s the history of the past half century of this field, so it should be no surprise that the terminology reflects this.
Read Gregory’s article for the short concise version of this dispute. What it means is that junk DNA threatens the worldviews of both Dembski and Dawkins!
Not quite. What you’re leaving out of this is the possibility of multiple levels of selection. In the original edition of The Selfish Gene (1976, p.76), Dawkins argued that “the simplest way to explain the surplus DNA is to suppose that it is a parasite, or at best a harmless but useless passenger, hitching a ride in the survival machines created by the other DNA”. Cavalier-Smith (1977) drew a similar conclusion (before he had read Dawkins), and Doolittle and Sapienza (1980) and Orgel and Crick (1980) [yes, that Crick] independently developed the concept of “selfish DNA” a few years later. This is an explicitly multi-level selection approach because it specifies that non-coding DNA can be present due to selection within the genome rather than exclusively on the organism (or gene, in Dawkins’s case) (see, e.g., Gregory 2004, 2005). (Incidentally, this idea of parasitic DNA dates back at least to 1945, when Gunnar Östergren characterized B chromosomes in this fashion). Of course, they tended to do what Ohno did and applied this one idea to all non-coding DNA, which is too ambitious. The modern view is more pluralistic (see, e.g., Pagel and Johnstone 1992 vs. Gregory 2003). Some non-coding DNA is just accumulated “junk” (in the definition of evidence-supported non-function that you espouse). Some (perhaps most) is “selfish” or “parasitic” and persists because there is selection within the genome as well as on organisms (in fact, an argument could be, and has been, made that “selfish DNA” would be a much more accurate term than “junk DNA” for most non-coding DNA). Some non-coding DNA is clearly functional at the organism level, including regulatory regions and chromosome structure components. Some of these latter functional non-coding DNA sequences are derived from elements that originally were of one of the first two types, most notably transposable elements that take on a regulatory function through co-option (or, in another manner of thinking, that undergo a shift in level of selection).
Junk DNA is not noncoding DNA and anyone who claims otherwise just doesn’t know what they’re talking about.
I’m afraid I don’t follow what you mean here. By your definition, “junk DNA” is any non-functional sequence of DNA, including pseudogenes (i.e., the original meaning). Those sequences do not encode proteins. Hence, your version of junk DNA is non-coding. I think this reflects the confusion that is imposed by the term “junk DNA”, which is why I generally think it is more obfuscating than enlightening.
________
References
Cavalier-Smith, T. 1977. Visualising jumping genes. Nature 270: 10-12.
Comings, D.E. 1972. The structure and function of chromatin. Advances in Human Genetics 3: 237-431.
Dawkins, R. 1976. The Selfish Gene. Oxford University Press, Oxford.
Doolittle, W.F. and C. Sapienza. 1980. Selfish genes, the phenotype paradigm and genome evolution. Nature 284: 601-603.
Gregory, T.R. 2003. Variation across amphibian species in the size of the nuclear genome supports a pluralistic, hierarchical approach to the C-value enigma. Biological Journal of the Linnean Society 79: 329-339.
Gregory, T.R. 2004. Macroevolution, hierarchy theory, and the C-value enigma. Paleobiology 30: 179-202.
Gregory, T.R. 2005. Macroevolution and the genome. In The Evolution of the Genome (ed. T.R. Gregory), pp. 679-729. Elsevier, San Diego.
Ohno, S. 1972. So much “junk” DNA in our genome. In Evolution of Genetic Systems (ed. H.H. Smith), pp. 366-370. Gordon and Breach, New York.
Orgel, L.E. and F.H.C. Crick. 1980. Selfish DNA: the ultimate parasite. Nature 284: 604-607.
Östergren, G. 1945. Parasitic nature of extra fragment chromosomes. Botaniska Notiser 2: 157-163.
Pagel, M. and R.A. Johnstone. 1992. Variation across species in the size of the nuclear genome supports the junk-DNA explanantion for the C-value paradox. Proceedings of the Royal Society of London, Series B: Biological Sciences 249: 119-124.